Credit: Wyss Institute at Harvard University
Fast, inexpensive test for malaria drills down to specifics
A team of researchers has developed a CRISPR-based malaria test that is fast, inexpensive and can be conducted in low-resource settings.
In ongoing efforts to eradicate malaria, an ultrasensitive test that offers rapid, species-specific diagnostic capabilities has been developed by a research collaboration led by James Collins , a core faculty member at Harvard’s Wyss Institute for Biologically Inspired Engineering.
To achieve the goal set by the World Health Organization’s (WHO) Global Malaria Control Programme , all local transmission of malaria parasites in defined geographic areas must be eliminated. Developing tests that are useful in the low-resource settings of many areas with endemic malaria is key.
Currently, the presence of the four major malaria-causing Plasmodium species P . falciparum , P . vivax , P . ovale , and P . malariae is determined by microscopic analysis of blood samples in which parasites can be detected in red blood cells, or with so-called rapid diagnostic tests for specific Plasmodium proteins (antigens).
“Unfortunately, available rapid diagnostic approaches cannot distinguish all four Plasmodium species from one other, which can be important to initiate the definitive course of treatment,” said Nira Pollock , associate medical director of Boston Children’s Hospital’s Infectious Diseases Diagnostic Laboratory and associate professor of pathology and medicine at Harvard Medical School (HMS). “And, most importantly, they are ineffective for detecting low numbers of Plasmodium parasites in asymptomatic individuals.”
“These ‘asymptomatic carriers’ are silent reservoirs for ongoing transmission by malaria-spreading mosquitoes and extremely important for ongoing global efforts to eradicate malaria,” added Jeffrey Dvorin , associate professor of pediatrics at HMS and senior associate physician in infectious diseases at Boston Children’s Hospital.
The research team, assembled by clinical fellow Rose Lee and included Pollock and Dvorin, created a field-applicable, ultrasensitive diagnostic assay that specifically detects DNA sequences from all Plasmodium species in symptomatic and asymptomatic malaria. The new method combines an optimized 10-minute rapid sample preparation protocol with the CRISPR-based SHERLOCK system to enable highly specific and sensitive Plasmodium detection in another 60 minutes in simple reporter devices. It is published in PNAS.
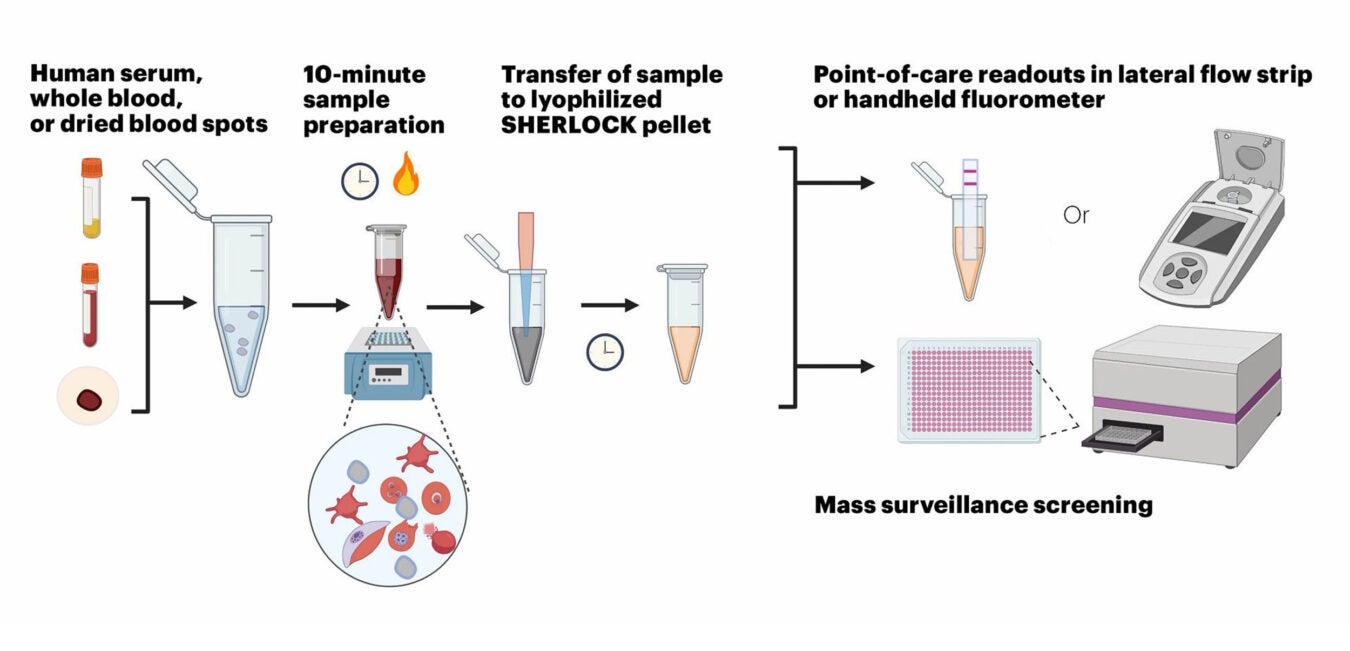
The Wyss Institute’s SHERLOCK malaria diagnostic test can diagnose specific Plasmodium parasites rapidly and ultra-sensitively in different types of samples, including human serum, whole blood, and dried blood spots.
Credit: Wyss Institute at Harvard University
The Wyss Institute’s SHERLOCK malaria diagnostic test can diagnose specific Plasmodium parasites rapidly and ultra-sensitively in different types of samples, including human serum, whole blood, and dried blood spots.
Credit: Wyss Institute at Harvard University
“This field-ready SHERLOCK diagnostic malaria assay surpasses the sensitivity and specificity requirements set by the WHO for a desired test that can be used to detect low parasite density in asymptomatic carriers of all major Plasmodium species,” said Collins. “Its highly streamlined design could provide a viable solution to the present diagnostic bottleneck on the path to eliminate malaria, and more generally enabling malaria surveillance in low-resource settings.”
Collins is a lead of the Wyss Institute’s Living Cellular Devices Focus Area, and also the Termeer Professor of Medical Engineering & Science at MIT.
Team members demonstrated their engineered SHERLOCK (specific high-sensitivity enzymatic reporter unLOCKing) assay to be capable of detecting less than two parasites per microliter of blood, the WHO’s suggested “limit of detection” (LOD) for a test with broad utility in endemic areas. Showing the assay’s clinical potential by analyzing clinical samples containing P . falciparum and P . vivax species, they called out them out with 100 percent sensitivity, by correctly identifying true positive samples, and 100 percent specificity, by also correctly identifying samples lacking a certain Plasmodium species in true negative samples.
Near 100 percent sensitivity and specificity are key attributes of diagnostic assays to be used in real-world testing. Moreover, the assay is designed so that it can also determine the presence of frequently mutated P . falciparum strains that have lost their HRP2 antigen and thus escape the detection by common rapid diagnostic tests.
Collins’ group at the Wyss Institute and MIT co-developed the SHERLOCK technology with Feng Zhang’s group at the Broad Institute. It was licensed to Sherlock Biosciences, a startup that used it to create a rapid molecular diagnostic for other disease applications, and recently received an emergency use authorization from the FDA for its COVID-19 rapid diagnostic.
Other methods have been developed that, like the new SHERLOCK assay, amplify and detect the DNA (or RNA) nucleic acid material of Plasmodium species. However, to date, these methods remain limited by their need for expensive laboratory equipment.
“Importantly, the assay is compatible with different sample types, such as whole blood, plasma, serum, and dried blood, and all components required for amplification, Cas12a activation and signal generation can be lyophilized in a single test tube to work together in a one-pot-reaction after they are reconstituted and mixed with patient sample,” said first-author Rose Lee, a clinical fellow in Collins’ group and Boston Children’s Hospital. “This … allows testing to be performed in low-resource settings with minimal expertise.”
Adapted from a Wyss Institute online article.
